Kiss 2012 Abstract Bioblast: Difference between revisions
Kiss Gergely (talk | contribs) No edit summary |
Gnaiger Caro (talk | contribs) No edit summary |
||
| (38 intermediate revisions by 6 users not shown) | |||
| Line 1: | Line 1: | ||
{{Abstract | {{Abstract | ||
|title=Kiss G (2012) | |title=Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C (2012) The negative impact of alpha-ketoglutarate dehydrogenase complex deficiency on matrix substrate-level phosphorylation. Mitochondr Physiol Network 17.12. | ||
|info=[[MiPNet17.12 Bioblast 2012|MiPNet17.12 Bioblast 2012 - Open Access]] | |info=[[MiPNet17.12 Bioblast 2012|MiPNet17.12 Bioblast 2012 - Open Access]] | ||
|authors=Kiss G | |authors=Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C | ||
|year=2012 | |year=2012 | ||
|event=[[Bioblast 2012]] | |event=[[Bioblast 2012]] | ||
|abstract= | |abstract=[[File:Kiss.G Bioblast2012 Abstract Photo(1).jpg|right|200px|Gergely Kiss]] | ||
Objectives: | '''Objectives:''' Provision of succinyl-CoA by the alpha-ketoglutarate dehydrogenase complex (KGDHC) is essential for generation of matrix ATP (or GTP) by substrate-level phosphorylation catalyzed by succinyl-CoA ligase. A decline in KGDHC activity has been associated with neurodegeneration. | ||
Methods: | '''Methods:''' Mitochondrial phosphorylation was investigated in tissues of transgenic mice with deficiencies in KGDHC subunits. | ||
Results: | '''Results:''' We demonstrate ATP consumption in respiration-impaired isolated and in situ neuronal somal mitochondria from transgenic mice with a deficiency of either dihydrolipoyl succinyltransferase (DLST) or dihydrolipoyl dehydrogenase (DLD) exhibiting a 20-48% decrease in KGDHC activity. Import of ATP into the matrix of mitochondria from transgenic mice was attributed to a shift in the reversal potential of the adenine nucleotide translocase towards more negative values due to diminished matrix substrate-level phosphorylation, causing the translocase to reverse prematurely. Immunoreactivity of all three subunits of succinyl-CoA ligase and maximal enzymatic activity were unaffected in transgenic mice as compared to wild-type littermates. Therefore, decreased matrix substrate-level phosphorylation was due to diminished provision of succinyl-CoA. These results were further corroborated by the finding that mitochondria from wild-type mice respiring on substrates supporting substrate-level phosphorylation exhibited ~30% higher ADP-ATP exchange rates compared to those obtained from DLST+/- or DLD+/- littermates. | ||
Conclusions: | '''Conclusions:''' We propose that KGDHC-associated pathologies are subserved by the inability of respiration-impaired mitochondria to rely on “in-house” mitochondrial ATP reserves. | ||
# [http://www.ncbi.nlm.nih.gov/pubmed/20207940 Chinopoulos C, Gerencser AA, Mandi M, Mathe K, Toeroecsik B, Doczi J, Turiak L, Kiss G, Konrad C, Vajda S, Vereczki V, Oh RJ, Adam-Vizi V (2010) Forward operation of adenine nucleotide translocase during F0F1-ATPase reversal: critical role of matrix substrate-level phosphorylation. FASEB J 24: 2405-2416. Open Access] | |||
|keywords=Succinyl-CoA ligase, Adenine nucleotide translocase, F<sub>0</sub>-F<sub>1</sub> ATP synthase, Reversal potential | |||
|mipnetlab=HU Budapest Chinopoulos C | |||
|journal=Mitochondr Physiol Network | |journal=Mitochondr Physiol Network | ||
|articletype=Abstract | |articletype=Abstract | ||
}} | }} | ||
{{Labeling|journal=Mitochondr Physiol Network | {{Labeling | ||
|area=Respiration, Genetic knockout;overexpression | |||
|organism=Mouse | |||
|tissues=Nervous system, Liver | |||
|preparations=Intact cells, Isolated mitochondria, Enzyme | |||
|enzymes=Adenine nucleotide translocase, Complex V;ATP synthase, TCA cycle and matrix dehydrogenases | |||
|injuries=Ischemia-reperfusion | |||
|diseases=Aging;senescence | |||
|topics=mt-Membrane potential, Substrate | |||
|instruments=Oxygraph-2k, O2k-Fluorometer | |||
|journal=Mitochondr Physiol Network | |||
|articletype=Abstract | |articletype=Abstract | ||
}} | }} | ||
| Line 24: | Line 38: | ||
== Affiliations and author contributions == | == Affiliations and author contributions == | ||
Gergely Kiss (1), Csaba Konrad (1), Judit Doczi (1), Anatoly A Starkov (2), Hibiki Kawamata (2), Giovanni Manfredi (2), Steven F Zhang (2), Gary E Gibson (3), M Flint Beal (2), Vera Adam-Vizi (1), Christos Chinopoulos (1,2) | |||
(1) Department of Medical Biochemistry, Semmelweis University, Budapest, 1094, Hungary; Email: [email protected] | |||
(2) Weill Medical College Cornell University, New York, NY, 10021, USA | |||
(3) Weill Cornell Medical College/Burke Medical Research Institute, White Plains, NY, 10605, USA | |||
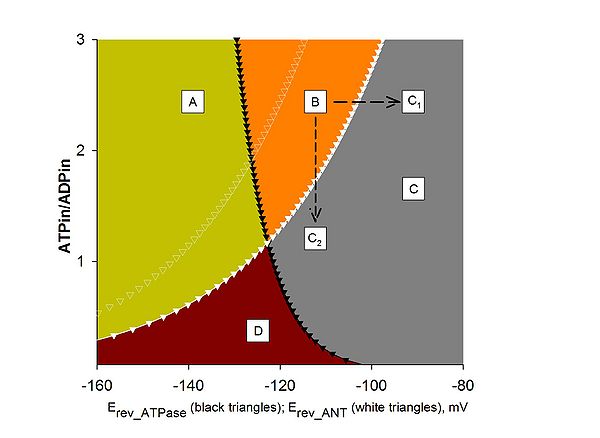
== Figure 1 == | |||
[[File:Kiss.G Bioblast2012 Abstract Figure.jpg|600px|Reversal potential of adenine nucleotide translocase]] | |||
Computational estimation of the reversal potential of adenine nucleotide translocase (Erev_ANT) and reversal potential of F<sub>0</sub>-F<sub>1</sub>ATPase (Erev_ATPase). A: ATPase forward, ANT forward; B: ATP reverse, ANT forward; C, C1, C2: ATPase reverse, ANT reverse; D: ATPase forward, ANT reverse. Black solid triangles represent Erev_ATPase; white solid triangles represent Erev_ANT. Values were computed for [ATP]out = 1.2 mM, [ADP]out = 10 μM, Pin = 0.01 M, n = 3.7 (2.7 plus 1 for the electrogenic ATP4-/ADP3- exchange of the ANT), pHi = 7.38, and pHo = 7.25. White open triangles represent Erev_ANT values computed | |||
for [ATP]out = 1.4 mM, and all other parameters as above. Traces have been computed by Erev estimator; the software and instructions on how to use it can be downloaded [http://www.oxphos.org/index.php?option=com_remository&Itemid=40&func=fileinfo&id=74 here]. | |||
== Help == | == Help == | ||
* [[MitoPedia: Terms and abbreviations]] | |||
* [[MitoPedia | |||
Latest revision as of 14:03, 16 June 2015
| Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C (2012) The negative impact of alpha-ketoglutarate dehydrogenase complex deficiency on matrix substrate-level phosphorylation. Mitochondr Physiol Network 17.12. |
Link: MiPNet17.12 Bioblast 2012 - Open Access
Kiss G, Konrad C, Doczi J, Starkov AA, Kawamata H, Manfredi G, Zhang SF, Gibson GE, Beal MF, Adam-Vizi V, Chinopoulos C (2012)
Event: Bioblast 2012
Objectives: Provision of succinyl-CoA by the alpha-ketoglutarate dehydrogenase complex (KGDHC) is essential for generation of matrix ATP (or GTP) by substrate-level phosphorylation catalyzed by succinyl-CoA ligase. A decline in KGDHC activity has been associated with neurodegeneration.
Methods: Mitochondrial phosphorylation was investigated in tissues of transgenic mice with deficiencies in KGDHC subunits.
Results: We demonstrate ATP consumption in respiration-impaired isolated and in situ neuronal somal mitochondria from transgenic mice with a deficiency of either dihydrolipoyl succinyltransferase (DLST) or dihydrolipoyl dehydrogenase (DLD) exhibiting a 20-48% decrease in KGDHC activity. Import of ATP into the matrix of mitochondria from transgenic mice was attributed to a shift in the reversal potential of the adenine nucleotide translocase towards more negative values due to diminished matrix substrate-level phosphorylation, causing the translocase to reverse prematurely. Immunoreactivity of all three subunits of succinyl-CoA ligase and maximal enzymatic activity were unaffected in transgenic mice as compared to wild-type littermates. Therefore, decreased matrix substrate-level phosphorylation was due to diminished provision of succinyl-CoA. These results were further corroborated by the finding that mitochondria from wild-type mice respiring on substrates supporting substrate-level phosphorylation exhibited ~30% higher ADP-ATP exchange rates compared to those obtained from DLST+/- or DLD+/- littermates.
Conclusions: We propose that KGDHC-associated pathologies are subserved by the inability of respiration-impaired mitochondria to rely on “in-house” mitochondrial ATP reserves.
• Keywords: Succinyl-CoA ligase, Adenine nucleotide translocase, F0-F1 ATP synthase, Reversal potential
• O2k-Network Lab: HU Budapest Chinopoulos C
Labels: MiParea: Respiration, Genetic knockout;overexpression Pathology: Aging;senescence Stress:Ischemia-reperfusion Organism: Mouse Tissue;cell: Nervous system, Liver Preparation: Intact cells, Isolated mitochondria, Enzyme Enzyme: Adenine nucleotide translocase, Complex V;ATP synthase, TCA cycle and matrix dehydrogenases Regulation: mt-Membrane potential, Substrate
HRR: Oxygraph-2k, O2k-Fluorometer
Affiliations and author contributions
Gergely Kiss (1), Csaba Konrad (1), Judit Doczi (1), Anatoly A Starkov (2), Hibiki Kawamata (2), Giovanni Manfredi (2), Steven F Zhang (2), Gary E Gibson (3), M Flint Beal (2), Vera Adam-Vizi (1), Christos Chinopoulos (1,2)
(1) Department of Medical Biochemistry, Semmelweis University, Budapest, 1094, Hungary; Email: [email protected]
(2) Weill Medical College Cornell University, New York, NY, 10021, USA
(3) Weill Cornell Medical College/Burke Medical Research Institute, White Plains, NY, 10605, USA
Figure 1
Computational estimation of the reversal potential of adenine nucleotide translocase (Erev_ANT) and reversal potential of F0-F1ATPase (Erev_ATPase). A: ATPase forward, ANT forward; B: ATP reverse, ANT forward; C, C1, C2: ATPase reverse, ANT reverse; D: ATPase forward, ANT reverse. Black solid triangles represent Erev_ATPase; white solid triangles represent Erev_ANT. Values were computed for [ATP]out = 1.2 mM, [ADP]out = 10 μM, Pin = 0.01 M, n = 3.7 (2.7 plus 1 for the electrogenic ATP4-/ADP3- exchange of the ANT), pHi = 7.38, and pHo = 7.25. White open triangles represent Erev_ANT values computed for [ATP]out = 1.4 mM, and all other parameters as above. Traces have been computed by Erev estimator; the software and instructions on how to use it can be downloaded here.